Introduction à l'anonymisation des données

Les données transmises à ARCHIMEDES doivent respecter la réglementation en vigueur en matière de protection de la vie privées et les autorisations éthiques. Dans de nombreux cas, cela implique d’anonymiser ou de coder les données avec le consentement des personnes concernées avant leur transmission.
Les outils et ressources ci-dessous sont fournis à titre purement informatif, et il incombe aux chercheurs de s’assurer que leurs données sont correctement préparées.
Le cycle de vie de l'anonymisation
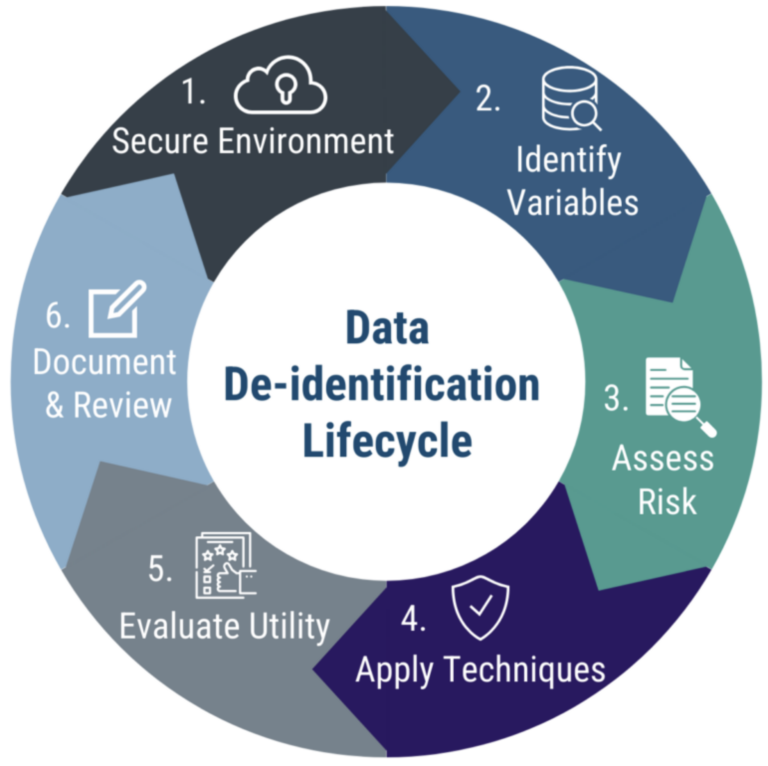
Le processus de pseudonymisation suit un cycle itératif. Chaque étape s’appuie sur la précédente pour limiter le risque de réidentification tout en préservant l’utilité des données.
- Environnement sécurisé — Effectuer la dépersonnalisation au sein d’un système sécurisé agréé.
- Identifier les variables — Identifier les variables directes et indirectes susceptibles de contribuer au risque de réidentification.
- Évaluer les risques – Évaluer la probabilité d’une réidentification en fonction du contexte et de l’utilisation prévue.
- Appliquer des techniques – Mettre en œuvre les méthodes appropriées pour réduire les risques identifiés en matière de confidentialité.
- Évaluer l’utilité — Évaluer l’impact sur la qualité des données et leur utilité analytique.
- Documenter et réévaluer – Consigner les décisions et procéder à une réévaluation périodique à mesure que le contexte évolue.
Les principes fondamentaux de l'anonymisation
Découvrez nos courtes vidéos de présentation générale qui abordent les principes fondamentaux de l’anonymisation des données de santé, la terminologie clé et les principaux facteurs de risque à prendre en compte.
Playlist

3:52

3:40
Welcome to ARCHIMEDES, the Advanced Research Collaboration for Health Integration, Medical Exploration, and Data Synthesis – a platform designed for seamless and secure medical data sharing.
Preparing data for sharing on ARCHIMEDES involves several steps to ensure privacy, security, and compliance with legal and ethical frameworks. One crucial component of this process is data de-identification. Data must be fully de-identified by the uploader before it is submitted to ARCHIMEDES.
De-identification is the process of removing or modifying personal information from data. This ensures that patient privacy is protected in medical data. It protects patient privacy, minimizes the risk of breaches, and allows data to be shared for collaboration – all while staying compliant with privacy laws.
However, de-identifying data isn’t always simple. It requires balancing privacy with data usability – and compliance with a range of regulatory frameworks. De-identification must account for potential risks of re-identification, especially with advances in data analytics and machine learning. Proper de-identification is essential for fostering trust in data sharing among stakeholders while preserving the value of the data for research and clinical use.
The terms “de-identification” and “anonymization” are often used interchangeably, but terminology can vary. Both processes remove personal health information (PHI) to protect privacy. Anonymization irreversibly removes PHI, which minimizes the risk of re-identification. On the other hand, de-identification (sometimes also called “pseudonymization”) removes most PHI, but may retain low-risk identifiers or use coding or encryption to preserve data utility over time. While both methods aim to protect privacy, de-identification often allows researchers to link data across time or datasets, whereas anonymization eliminates this possibility for greater privacy protection. Both anonymization and de-identification protect privacy and ensure compliance with privacy regulations, but de-identification often allows for greater data utility. To achieve this, a variety of techniques can be used to effectively remove or alter sensitive information. Let’s explore some of the most commonly used de-identification methods
First, data masking. This involves the removal or modification of direct identifiers—things like names, phone numbers, and medical record numbers. Masking is often the first and most straightforward step in the de-identification process.
Next, data perturbation. This method slightly modifies the values of sensitive data to protect identity. For example, an age or date might be adjusted by a small, random amount. While the overall dataset stays statistically meaningful, individual-level precision is blurred to protect privacy.
Finally, tokenization. This replaces identifiable data with unique codes or pseudonyms that cannot be linked back to an individual without a secure key. Tokenization is especially helpful when researchers need to track records across time or across datasets, without compromising identity.
Together, these tools form the foundation of most de-identification strategies—removing identifiers, adding uncertainty, and preserving utility where possible.
In Canada, the Personal Information Protection and Electronic Documents Act—PIPEDA— outlines legal requirements for de-identifying medical data. In the U.S., the HIPAA De-identification Standard outlines similar rules. These frameworks define how data must be treated. While the PIPEDA outlines legal requirements for data de-identification, the Office of the Privacy Commissioner of Canada provides guidance on how to adequately de-identify data. Some provinces also have their own regulations. There are lots of other resources available to learn more about de-identification regulations.
To explore resources, templates, and tools for data de-identification, visit the ARCHIMEDES platform and learn how to get started.
Welcome to Archimedes, the advanced research collaboration for health integration, medical exploration, and data synthesis. A platform designed for seamless and secure health data sharing.
Different types of data require different deidentification methods. Here we provide a basic intro to some high-level considerations for different types of medical data. Let’s start with considering deidentification of tabular data. Typically, this type of data includes patient demographics, survey scale responses, treatment records, and clinical notes. To approach deidentifying tabular data, you are going to need to remove direct identifiers, aggregate sensitive fields, or engage in data perturbation to ensure compliance with HIPAA guidelines.
Next, we will talk about deidentification approaches for clinical data. Clinical data may include patient records from hospital systems or electronic health records or EHRs which often contain structured and unstructured information such as lab test results, prescriptions, clinical notes, and diagnostic codes. Deidentifying this data requires masking of obvious identifiers, generalizing or binning sensitive numerical fields like age or weight, and careful handling of free text notes to remove contextual identifiers. Natural language processing tools may be used to help scrub unstructured notes while maintaining meaning.
Lastly, we will talk about the approach for deidentifying imaging data. Imaging data may include radiology images such as ultrasound scans, MRI, CT, etc. Most of these images are saved in the standard DICOM format. DICOM imaging data contains the visible image as well as a text component called metadata which contains information about the image such as the kind of machine that was used to take the image, the date or time when the image was taken, patient information such as name and ID number and more.
The metadata text component that is attached to the image is not seen visually but can be accessed using viewing software and or programming languages. Deidentification of DICOM metadata follows the same approach as tabular data that is clearance of certain PHI including blanking of certain tags and or tokenization of others etc. This covers the approach for deidentification of the metadata.
But what about the image itself? Careful. Some imaging data include burned-in PHI in the pixels of the image itself. This is a common feature of ultrasound images. In these cases the pixel data also needs to be deidentified. Approaches to this could include applying blurring, pixelation, or blackboxing to overlay and therefore hide the PHI. Alternatively, a cropping approach could be used to mask patient features. Optical character recognition (OCR) could also be used to recognize and black out text to anonymize any associated labels, or it can be used in QA or validation workflows to find PHI in images before deidentification.
This flowchart summarizes some examples of deidentification steps by data type. Use it as a guide to determine the right approach for your data set and always confirm that your process meets the legal and institutional requirements for ethical data sharing.
Coming up next, we’ll walk you through detailed tutorials on how to deidentify different types of data using free and open-source tools. We’ve just covered the foundational concepts of deidentification, but theory is only the first step. In the next videos, we’ll walk through hands-on tutorials that show how to apply these techniques using real world examples and free tools. Whether you’re working with spreadsheets or DICOM files, to explore resources, templates, and tools for data deidentification, visit the Archimedes platform and learn how to get started.
Vois aussi
- Processus de pseudonymisation (à venir)
- Tutoriels et ateliers (à venir)
- Bibliothèque de liens (à venir)
